 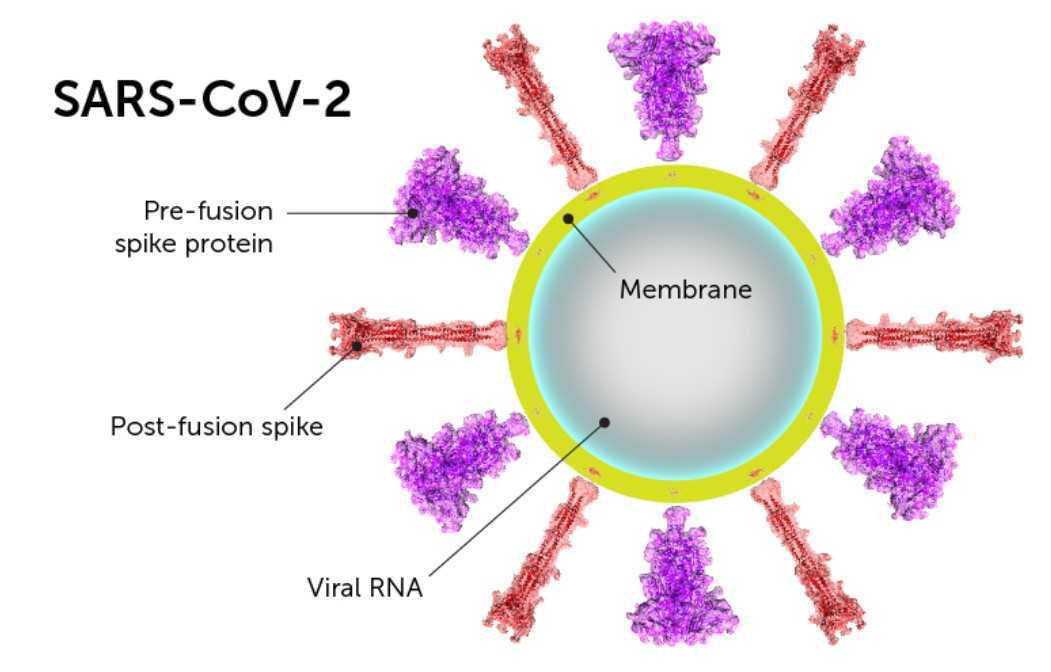
We found that about 50% of the mutability maps variability of the S protein of the severe acute respiratory syndrome-related betacoronavirus (sarbecovirus), and 67% of the variability in the mutability of the seasonal influenza spike (HA) can be attributed Ab pressure, as estimated from the model. Next, we developed a computational approach based on spectral clustering to compare these maps. We then used sequences from public repositories ( to evaluate the mutability maps of those glycoproteins. Superimposed on the spike surface, the immunogenicity score gives the Ab affinity maps of influenza and corona spikes, which we applied to predict how the antigenic space is explored unevenly across the surface of these glycoproteins. We developed here an in-silico approach to estimate the Ab targeting - a proxy for B cell immunogenicity ( 24), of residues on the spike surface, and the differential accessibility to antigenic epitopes due to the geometrical presentation of spikes on the surface of the virus. We have previously shown using coarse-grained molecular dynamics simulations, that the geometry of the immunogen spike presentation on the virus recapitulates the known immunodominance of hemagglutinin (HA) head compared to its stem ( 24). Here, we concentrate on the geometric presentation of the spike to Abs. Amongst them is the B cell repertoire - the number of B cell clones targeting different epitopes ( 24 – 29), their germline affinity ( 24, 30), and T cell help to B cell ( 31). The magnitude (titers) of Ab response against a given epitope is a direct consequence of the B immunodominance hierarchy patterns of an immunogen, which are the result of various aspects of the humoral response to antigen ( 21 – 23). Here we sought to understand and predict, from first principle, to what extent the mutability of the spikes of influenza and close relatives of SARS-CoV-2 could be attributed to Ab pressure. More generally, antigenic drift due to Ab pressure is common in other RNA viruses such as the seasonal influenza virus ( 19, 20). Hence, given the prevalence of SARS-CoV-2, to inform vaccine design and understand how the fitness landscape of the virus evolves, it is important to recognize antigenic drift due to Ab pressure if it were to occur. However, other members of the coronavirus family have been circulating in human populations for many years ( 18) and evidence of antigenic drift is seen in SARS-CoV-1 (Guan et al., 2003 Song et al., 2005), and among common cold coronaviruses 229E (Chibo and Birch, 2006). One mutation at the spike (D614G) is now widespread and is thought to support a high viral growth rate ( 17). Since SARS-CoV-2 virus introduction into humans is recent, it probably has not yet evolved extensively to acquire escape mutations from the commutative Ab pressure of the human population ( 16). In particular, Abs against the spike receptor-binding domain (RBD) have been shown to have neutralization and protective capabilities ( 14, 15). These therapeutic approaches, hopefully, would be able to elicit strong Ab and T cell response against the virus. The spike, a class I fusion protein, mediates entry to the host cell by binding to the angiotensin-converting enzyme 2 (ACE2) receptor (Ou et al., 2020) and is the main target of Ab response (Robbiani et al., 2020 Wu et al., 2020). Almost all vaccination approaches aim to use the glycoproteins or spike protein (S) of the virus in its trimeric form ( 12) or vaccinate with the full (inactivated) virus ( 13). In response to the SARS-CoV-2 pandemic, many approaches for antibody (Ab) therapies, and vaccines are being explored ( 11). One result of such proofreading activity is that CoVs genomes are less mutable compared to other RNA viruses ( 9), and thus the sequence diversity of SARS-CoV-2 is quite low ( 10). Nonstructural protein 14 (nsp14), a subunit of the replicase polyprotein encoded by CoVs is thought to provide a form of proofreading activity that could support the expansion of large CoVs genomes to their current size. It is a member of the betacoronaviruses family ( 4), related to coronaviruses found in Bats ( 5), and to SARS CoV that cause severe respiratory syndrome ( 6) as well as other widely circulating members of the family that cause the common cold ( 7).Ĭoronaviruses (CoVs) have the largest genomes among RNA viruses ( 8). The virus emerged as a result of a zoonotic shift ( 2, 3). 
The COVID-19 pandemic, caused by the SARS-CoV-2 coronavirus, is one of the most challenging global health crisis within the last century ( 1). 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
